RFP Lab
Purpose - To make RFP from jelly fish in bacteria, and to learn about steps in genetic engineering.
Materials and Procedure -
Lab: 2a- materials and procedure can be found in Amgen lab manual part 2a
4a -materials and procedure can be found in Amgen lab manual part 4a
5a - materials and procedure can be found in Amgen lab manual part 5a
6a - materials and procedure can be found in Amgen lab manual part 6a
Experimental Overview -
2a: We verified that our DNA had the plasmid we wanted by using restriction digest, and then we used BamH1 and Hindi III to cut the RFP-ara from bacterial plasmid.
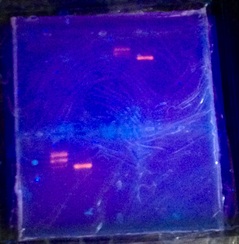
4a: We verified that we had the plasmid digest though electrophoresis.
5a: We used a recombinant plasmid to transform the bacteria.
6a: We then purified our RFP using chromatography.
Results -
Questions -
2a Pre-lab:
1. Two fragments are made, one is the RFP with PBAD that is 807 BP, and one is Ara- combined with ori combined with Amp-R that is 4495 BP.
2. The RFP and Ara-C are required.
3. The selectable marker allows the separation of bacteria that doesn't have the desired gene.
2a:
1. Ori= origin of replication; RFP= gene of interest; Amp-R = antibiotic resistance and selective marker; Ara-C = binds to promoter
2. The resistance enzymes cut up unwanted bacteria DNA.
3. The bacteria keeps antibiotic resistant genes in order to survive. In modern medicine new stronger antibodies are created to kill bacteria as old ones no longer work.
4. Central dogma is the same for all organisms.
5. Take half of the mixture and put each half in a petri dish. Then, put Amp in the Kan dish to kill Kan. Then in put Kan in the Amp dish to kill Amp. This separates Amp and Kan from each other.
4a:
1. We make ampicillin resistant bacteria so that cells with Amp-R genes will live.
2. The pARA-R plasmid will only turn on the promoter if the bacterial cells are given arabinose. Without arabinose, then they won't turn red.
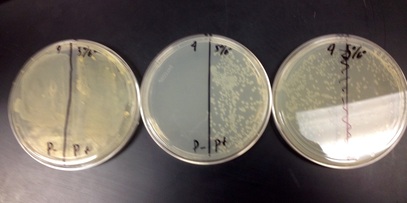
3. P- and P+ will grow on the LB plate, only P+ will grow on the LB/Amp plate, and a small amount of bacteria will grow on the LB/Amp/Ara plate.
Materials and Procedure -
Lab: 2a- materials and procedure can be found in Amgen lab manual part 2a
4a -materials and procedure can be found in Amgen lab manual part 4a
5a - materials and procedure can be found in Amgen lab manual part 5a
6a - materials and procedure can be found in Amgen lab manual part 6a
Experimental Overview -
2a: We verified that our DNA had the plasmid we wanted by using restriction digest, and then we used BamH1 and Hindi III to cut the RFP-ara from bacterial plasmid.
4a: We verified that we had the plasmid digest though electrophoresis.
5a: We used a recombinant plasmid to transform the bacteria.
6a: We then purified our RFP using chromatography.
Results -
Questions -
2a Pre-lab:
1. Two fragments are made, one is the RFP with PBAD that is 807 BP, and one is Ara- combined with ori combined with Amp-R that is 4495 BP.
2. The RFP and Ara-C are required.
3. The selectable marker allows the separation of bacteria that doesn't have the desired gene.
2a:
1. Ori= origin of replication; RFP= gene of interest; Amp-R = antibiotic resistance and selective marker; Ara-C = binds to promoter
2. The resistance enzymes cut up unwanted bacteria DNA.
3. The bacteria keeps antibiotic resistant genes in order to survive. In modern medicine new stronger antibodies are created to kill bacteria as old ones no longer work.
4. Central dogma is the same for all organisms.
5. Take half of the mixture and put each half in a petri dish. Then, put Amp in the Kan dish to kill Kan. Then in put Kan in the Amp dish to kill Amp. This separates Amp and Kan from each other.
4a:
1. We make ampicillin resistant bacteria so that cells with Amp-R genes will live.
2. The pARA-R plasmid will only turn on the promoter if the bacterial cells are given arabinose. Without arabinose, then they won't turn red.
3. P- and P+ will grow on the LB plate, only P+ will grow on the LB/Amp plate, and a small amount of bacteria will grow on the LB/Amp/Ara plate.
5a:
1. Yes, our results were correct because we had growth on the LB plate for P+ and P-, the LB/Amp plate only grew on the positive side, and the LB/Amp/Ara plate had growth.
2. Their were no red colonies apparent most likely from temperature or not enough time staying in the incubator.
3. The RFP can only appear on the LB/Amp/Ara plate because RFP needs arabinose to be expressed.
4. The more copies apparent, then the more likely the promoter will be on and visible.
5. The RFP gene is shown from a trait through transcription by DNA --> mRNA --> Protien (process of central dogma).
6. Since central dogma is the same in all organisms, bacteria can replicate human genes.
1. Yes, our results were correct because we had growth on the LB plate for P+ and P-, the LB/Amp plate only grew on the positive side, and the LB/Amp/Ara plate had growth.
2. Their were no red colonies apparent most likely from temperature or not enough time staying in the incubator.
3. The RFP can only appear on the LB/Amp/Ara plate because RFP needs arabinose to be expressed.
4. The more copies apparent, then the more likely the promoter will be on and visible.
5. The RFP gene is shown from a trait through transcription by DNA --> mRNA --> Protien (process of central dogma).
6. Since central dogma is the same in all organisms, bacteria can replicate human genes.

6a:
1. The red florescent protein (RFP) can be seen separated because the cells are red and florescent.
2. The supernatant was fairly clear and the pellet had a pink or light red tint.
6b:
1. BB (Binding Buffer) - makes amino acid and protein bind to the resin beads.
WB (Wash Buffer) - removes the proteins not bound to the resin beads
EB (Elution Buffer) - takes protein off resin beads
CEB (Column Equilibration Buffer) - stores resin beads
2. This time the supernatant was more of a see through pink and the pellet was more of a dark pink/red color.
Analysis - Our lab was successful. We knew that the bacteria was present from our gel electrophoresis, so we needed to know if we had successfully grown it and could isolate it. When we put the wash buffer through the column, we could see the red proteins on the beads.
UPDATE: We ran the protein through gel electrophoresis to check for purity and size. We should have seen a strong, single band at the 20-25 mark, but instead we saw a faint band at the 15-20 mark. This indicates that we had either a different protein, it degraded, and/or part of it was clipped off.
Reflection - I had mixed feelings about this lab. It was verrrrrrrry long, almost frustratingly long. However, it was pretty satisfying to finally see those little red proteins, and we were able to see how we could transform and isolate bacteria. That was the best part about this experiment, seeing how the central dogma being universal helps us transfer genes between humans and bacteria. As far as my group is concerned, we had a good group. No one slacked off, we never argued, and we even had a few fun/funny moments. While this project was long and sometimes boring, I was happy with the end result.
1. The red florescent protein (RFP) can be seen separated because the cells are red and florescent.
2. The supernatant was fairly clear and the pellet had a pink or light red tint.
6b:
1. BB (Binding Buffer) - makes amino acid and protein bind to the resin beads.
WB (Wash Buffer) - removes the proteins not bound to the resin beads
EB (Elution Buffer) - takes protein off resin beads
CEB (Column Equilibration Buffer) - stores resin beads
2. This time the supernatant was more of a see through pink and the pellet was more of a dark pink/red color.
Analysis - Our lab was successful. We knew that the bacteria was present from our gel electrophoresis, so we needed to know if we had successfully grown it and could isolate it. When we put the wash buffer through the column, we could see the red proteins on the beads.
UPDATE: We ran the protein through gel electrophoresis to check for purity and size. We should have seen a strong, single band at the 20-25 mark, but instead we saw a faint band at the 15-20 mark. This indicates that we had either a different protein, it degraded, and/or part of it was clipped off.
Reflection - I had mixed feelings about this lab. It was verrrrrrrry long, almost frustratingly long. However, it was pretty satisfying to finally see those little red proteins, and we were able to see how we could transform and isolate bacteria. That was the best part about this experiment, seeing how the central dogma being universal helps us transfer genes between humans and bacteria. As far as my group is concerned, we had a good group. No one slacked off, we never argued, and we even had a few fun/funny moments. While this project was long and sometimes boring, I was happy with the end result.